The Endless Battle Between Viruses & Bacteria
If you haven’t heard, CRISPR is easily one of the biggest and most important scientific stories over the past decade. It is a simple yet powerful tool for researches to edit our genes. That’s right, the ability to alter our DNA and modifying genetic functions is no longer a science-fiction fantasy. Among the potential applications include correcting any genetic defects, treatment and prevention of diseases and improving our food crops.
What’s interesting to note is that the technology was actually derived from the natural defence mechanisms of bacteria and archaea (single-celled microorganisms or prokaryotes). These organisms developed a specific adaptive immune system against the virus.
Let’s take a closer a look to understand the origins of CRISPR.
Drivers of Evolution
The evolutionary cycle of our planet’s countless organisms is fascinating. The story behind it however, can be ruthless. Only the strong can survive and they continue to expand, while the weaker ones eventually becomes extinct. In the case of fungi, bacteria and viruses; billions of these microorganisms are at war every day, all around us. They fight and kill each other in environments where nutrients are limited.
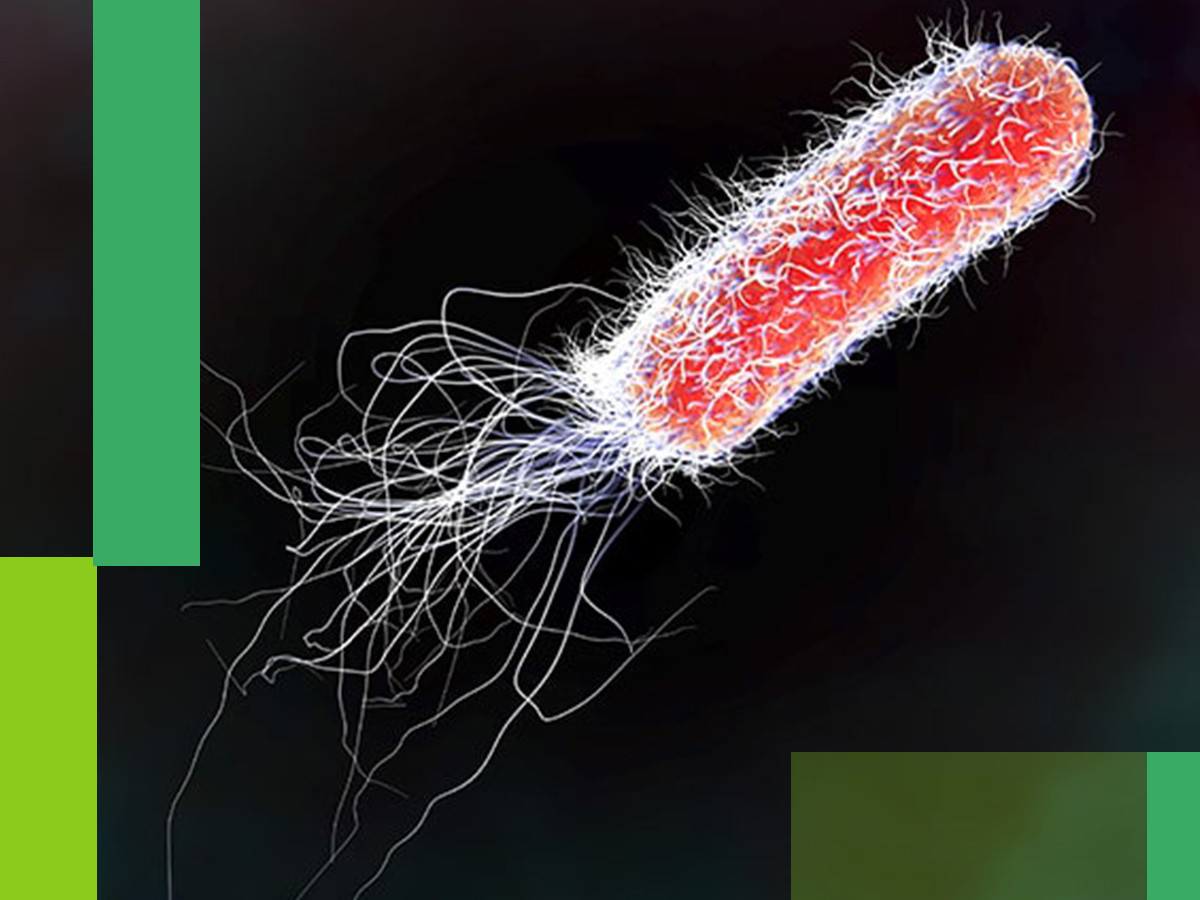
Every predator is prey to something else. While the bacteria can infect even the largest and strongest of organisms (us humans included); even they fall prey to the virus. Just as there are viruses infecting human cells, there are viruses known as bacteriophage that prey on bacteria.
Phages Outnumber All Living Organisms
The deadliest being on earth is undoubtedly the phage virus. There are more phages on earth than all other living organisms combined. Every day, trillions of bacteria are killed due to bacteriophage infection. This includes up to 40% of ocean bacteria, every day.
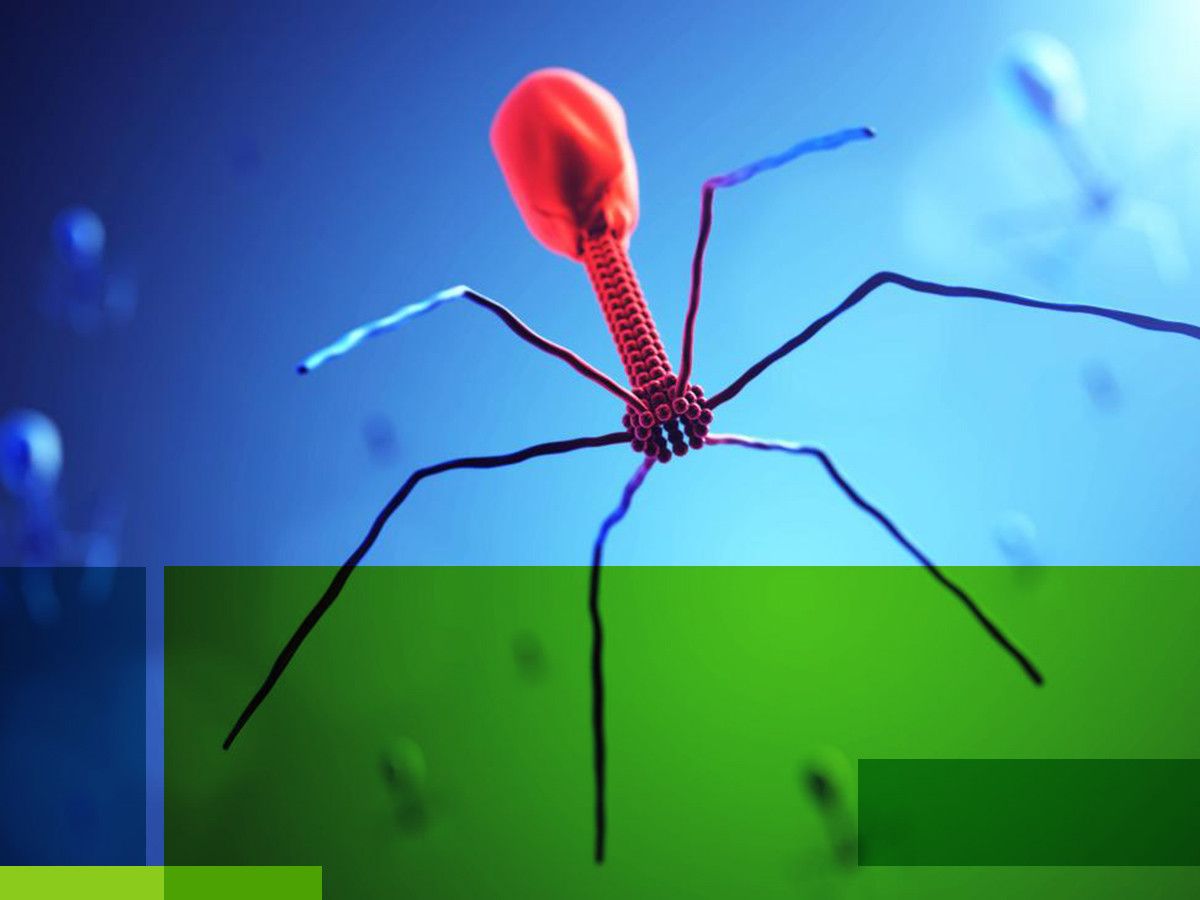

It is important to note that viruses are widely regarded as non-living entities. They are typically made of a core genetic code (DNA/RNA) which is encased within a protein coat called a capsid. Most viruses have lipids surrounding the capsid in an envelope. In the case of the bacteriophage, the infamous structure of the T4 looks unnatural. It is non-enveloped, has a contractile tail and fibres to latch onto bacteria cell walls.
The Vicious Cycle
Now, viruses cannot replicate on their own. Instead, they infect hosts that are deemed compatible. Upon latching onto a bacteria’s cell wall, the spike at the phage’s tail end ‘injects’ the viral DNA into the bacteria, hijacking the bacteria’s cellular functions. The usual processes for producing bacterial components for replication are now taken over to produce the components of the virus. These components are then assembled into new bacteriophage offspring.

Within the bacteria cell wall, all known resources are consumed until it is filled with new bacteriophages. As it reaches maximum capacity, the phages break free, killing its bacterial host. This is known as the ‘lytic’ cycle.
In a ‘lysogenic’ cycle, the infection is more methodical. The injected phage DNA first integrates into the bacterial chromosome to produce the ‘prophage’. When the bacterium reproduces or multiplies, the prophage is also copied and is present in each of the bacterial daughter cells. These daughter cells can continue to replicate with the prophage present; or the prophage can exit the bacterial chromosome to initiate the lytic cycle.
What Doesn’t Kill You…
So, here’s where evolution kicks in. While a lot of bacteria are killed in what seems to be an eternal vicious cycle, some do survive. They developed an elegant defence mechanism known as CRISPR-Cas.
Let’s go back to the year 1987, when Japanese scientists were studying the popular bacteria E. coli. They documented repeating sequences in the bacteria’s DNA, stating them to have unknown biological significance. Over the years, other researchers found similar clusters in the DNA of other bacteria (and archaea). They eventually gave these sequences a name: Clustered Regularly Interspaced Short Palindromic Repeats — or CRISPR.
Then again, these sequences were regarded as a mystery until 2007. At the time, food scientists were studying the Streptococcus bacteria used to make yogurt, and they discovered something remarkable. These odd clusters serve an important function; as part of the bacteria’s immune system.
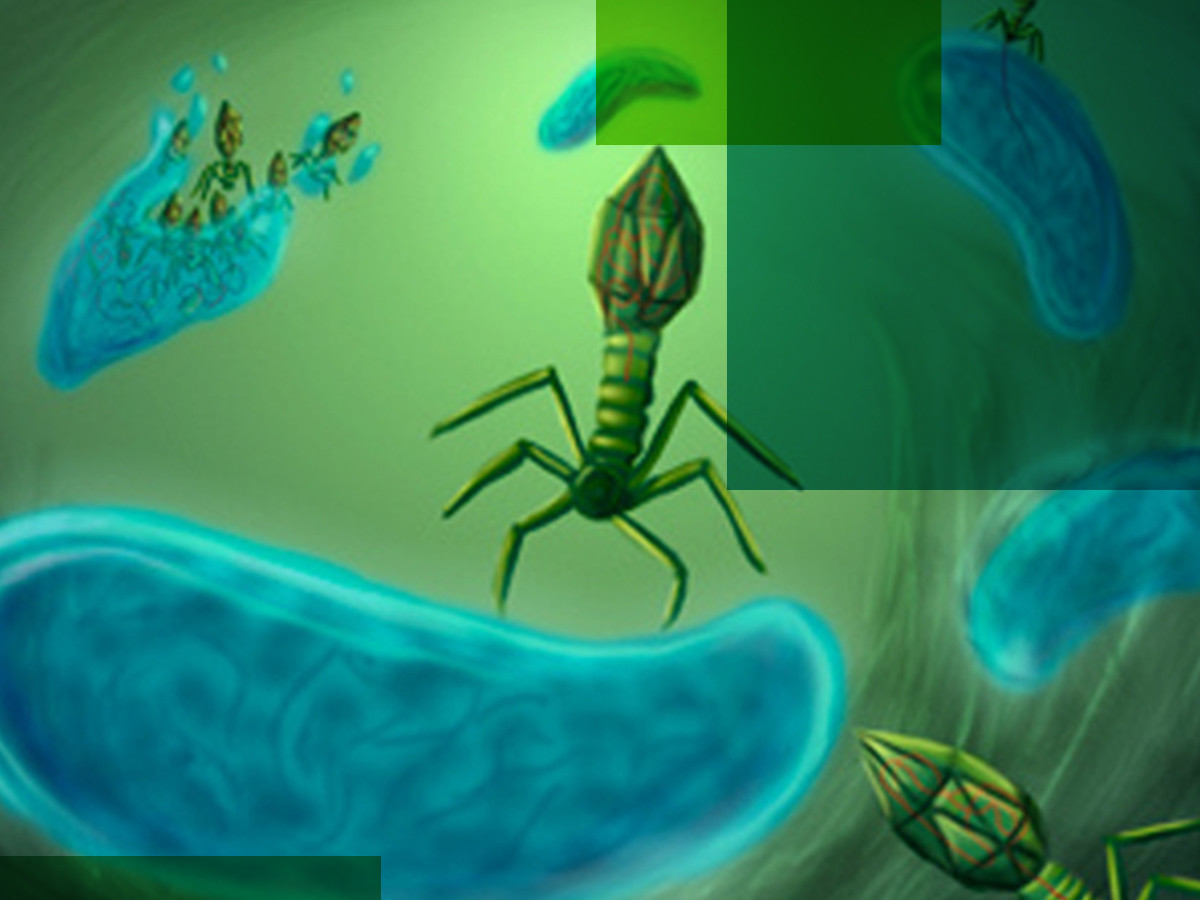
Bacteria are under constant assault from viruses. Consequently, they produce specific enzymes to fight off viral infections. Whenever those enzymes kill off an invading virus, other smaller enzymes will come along and pick up the remains of the virus’s genetic code. They acquire these codes by cutting it into bits, and then storing them into the bacteria’s CRISPR spaces.
In short, the bacteria are able to acquire and permanently maintain a record of the information carried in the spacer sequence.
…Makes You Smarter
Here’s the clever part of this elegant defence mechanism: the bacteria use the genetic information stored in these CRISPR spaces to defend themselves from future attacks. Whenever a new virus attacks, the bacteria produce special enzymes, known as Cas9 (CRISPR associated protein 9), that carry around those stored bits of viral genetic code like a wanted poster or suspect ID. When these Cas9 enzymes come across a virus, it scans the virus’s DNA against the expressed RNA (transcribed from the CRISPR sequences) to see if they match. If there’s a match, the Cas9 enzyme starts chopping up the virus’s DNA to neutralise the threat.
What makes the defence mechanism more robust is its ability to store different sequences. As the name implies, CRISPR refers to the short, repetitive sequences that separate each unique spacer sequence. This means that different ‘spacers’ can come from several different phages, therefore each bacterium has a unique sequence stored in its genetic database.
Practical Applications Of CRISPR
There are several diverse applications where we can immediately put CRISPR to good use. Since they are unique, spacers can be used to identify the bacteria that carry them.
Even for something as seemingly ordinary as the dairy industry, CRISPR has a place – for cheese production. We all know that the process of making cheese requires very high densities of bacterial cultures to keep up with the high volume of production. This makes them extremely vulnerable to the threat of phage infections, which can kill the entire bacterial population while leaving a lot of phages in the product. By introducing CRISPR sequences into these bacteria, we can make them resistant to phage infections.
One common infection that we humans must deal with is Salmonella, a bacterial disease that affects the intestinal tract. Although current methods for detecting Salmonella infections are accurate, they are technically demanding, thus making it difficult to implement in poorer countries. Additionally, these methods are non-automated, which delays detection and further presents a bottleneck. However, identification of ‘signature’ CRISPR sequences can be automated, thus helping detect bacterial pathogens and thereby the corresponding bacterial subtype, quickly and accurately.
Conclusion
The constant battle of life or death between viruses and bacteria has raged for millions of years. It will continue all around us, every day. As scientists, researchers and innovators however, it’s a war that we humans can study, adapt and effectively use to our advantage. Implemented correctly, CRISPR-Cas technology can be used to enhance human life; from food production to healthcare.
For more information, contact us at xeraya@xeraya.com or visit our website www.xeraya.com
Sources:
- https://www.sciencedirect.com/science/article/pii/S030090841500104https://www.livescience.com/58790-crispr-explained.html
- https://www.vox.com/2018/7/23/17594864/crispr-cas9-gene-editing
- https://thewire.in/the-sciences/how-a-battleground-between-bacteria-and-viruses-is-being-used-for-human-health
- https://www.livescience.com/27098-ocean-viruses-battle-bacteria.html
- https://misciwriters.com/2016/04/19/virus-vs-bacteria-mortal-combat/




